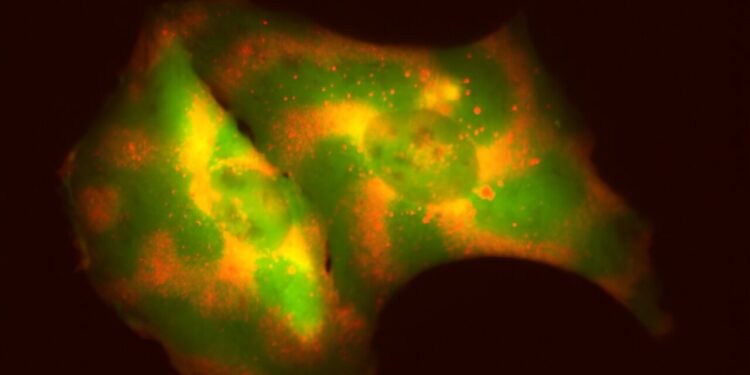
The Coyle lab tested its tool on human cells by programming several proteins, each represented by a different color, to move in different wave patterns. The tool they developed allows researchers to essentially program molecules to move within a cell to specific locations over time. Credit: University of Wisconsin-Madison
Researchers can engineer cells to express new genes and produce specific proteins, giving the cells new things to work with. But it is much more difficult to provide cells with instructions on how to organize and use these new parts. Now, new tools developed by researchers at the University of Wisconsin-Madison offer an innovative solution to this problem.
Their research is published in the journal Cell.
Everything a cell does depends on how molecules are organized within the cell. Inside our cells – all cells – proteins and other molecules undergo organization and reorganization to carry out cellular function. Like a fleet of commuter trains moving at regular intervals along their various routes, a cell’s proteins are organized in time and space to perform complex but predictable functions.
Although the need to organize molecules within a cell is universal among living organisms, the specific proteins and mechanisms responsible for this organization vary. In a system specific to bacterial cells, for example, the proteins MinD and MinE, known collectively as MinDE, interact with each other along the cell membrane to produce wave patterns that facilitate the movement of molecules within the cell.
When molecules fail to organize properly within a cell, it can have serious consequences, including uneven cell division and poor communication within and between cells, both of which are associated with developmental disorders. and diseases such as cancer.
In short, we know how to add new parts to cells, but it is much more difficult to provide instructions on how to organize and use them.
The mechanisms by which molecules organize themselves and interact with each other have been refined over millennia of evolution. When scientists engineer cells to produce new molecules, it is difficult to get the cells to use these new molecules without unintentionally disrupting other natural cellular functions.
UW-Madison biochemists have developed a tool to control the movement and organization of specific proteins in mammalian cells while leaving other proteins alone. Their new tool exploits waves and oscillations derived from interactions between MinDE proteins, which are found only in bacteria and do not interfere with mammalian cellular function.
By engineering interactions between MinDE proteins and proteins of interest, researchers created highly specified models to organize molecules within mammalian cells and induce cellular behaviors and functions. The tool allows researchers to adjust and change patterns in response to stimuli, essentially programming molecules to move within a cell to specific locations over time.
This innovative tool has multiple potential uses for scientists interested in engineering specific cellular activities or studying cellular activity in a living organism.
Controlling the ratio of MinDE proteins allows researchers to design movement patterns that determine how molecules are organized in a cell, which could be used to orchestrate cellular activities such as moving or communicating with other cells .
Variation in movement patterns can also be used to study cellular activity. As each MinDE protein report emits a unique oscillatory pattern, proteins can be inserted into a group of cells to give each cell its own pattern – an individual beacon that allows researchers to observe each cell more easily.
Scientists can also use these unique models to analyze cell signaling patterns to learn the shape, location, and signaling activity of each individual cell. Researchers at UW-Madison’s Coyle Lab liken this use of the tool to an FM radio dial, because it allows them to listen to the unique data emitted by each cell in a multi-cell system, typically a very difficult task.
The Coyle lab plans to continue exploring applications of the tool, including studying the dynamics of signaling pathways in tumors, a classic example of impaired cellular behavior and function.
More information:
Rohith Rajasekaran et al, A programmable reaction-diffusion system for designing spatio-temporal cellular signaling circuits, Cell (2024). DOI: 10.1016/j.cell.2023.12.007
Provided by University of Wisconsin-Madison
Quote: Programming cells to organize their molecules could open the door to new treatments (February 15, 2024) retrieved February 16, 2024 from
This document is subject to copyright. Except for fair use for private study or research purposes, no part may be reproduced without written permission. The content is provided for information only.