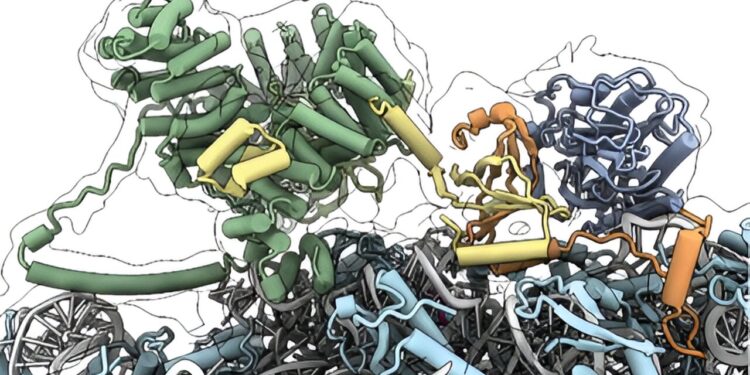
Molecular model of the cotranslational multi-enzyme complex comprising the ribosome (blue and gray), the NAC chaperone complex (yellow and orange), and the enzymes MetAP1 (cambria) and NatA (green). Credit: Denis Yudan and Nenad Ban
Proteins are among the most important molecular building blocks of life. They themselves consist of amino acids linked together based on information contained in our genetic material. In this process, the genetic code is translated into an amino acid sequence. However, this translation is only the first step. Often, special enzymes modify the new (nascent) proteins as they leave their cellular production site, the ribosome. Only then can the proteins fulfil their various biological functions.
Until now, it was not well understood how these enzymes interact to modify nascent proteins at the ribosome, nor how their activity is regulated and coordinated. Now, an international team of researchers from Caltech, the University of Konstanz and ETH Zurich has elucidated the complex molecular mechanism of two consecutive protein modifications that affect about 40% of all mammalian proteins. It turns out that the star of the show, a kind of molecular control center, is a relatively small chaperone protein complex called NAC (nascent polypeptide-associated complex).
Without NAC, all sorts of stress responses are triggered, causing proteins to misfold or be sent to the wrong places in the body.
“The essential chaperone NAC has been implicated or involved in a myriad of processes. It has been difficult to understand why a single small protein complex that seems very simple would impact all these different processes,” says Shu-ou Shan, Altair Professor of Chemistry at Caltech, executive director of biochemistry and molecular biophysics and corresponding author of a new paper describing the work.
“But we are now beginning to see the full picture of NAC as a master higher-order regulator of protein production in the cell. It has become clear that the job of NAC is to recruit various biogenesis factors to the ribosome and associate them with the appropriate nascent proteins that are being synthesized.”
The results are published in the journal Nature.
Essential for normal cell functioning
In the new paper, the researchers focus on two modifications orchestrated by NAC: the removal of the first amino acid, methionine, from the growing protein, followed by the attachment of what’s called an acetyl group to the remaining end.
Both of these processes take place at the exit of the ribosomal tunnel, i.e. where proteins leave the ribosome as a growing chain of amino acids during synthesis. These modifications are essential for the majority of our proteins, as they influence multiple properties of the proteins, such as their three-dimensional folding, their lifespan or their interaction with other proteins, and therefore the proper functioning of the proteins.
“Dysregulation of the processes involved in protein modification can have extremely negative consequences for the organism. It is associated with developmental disorders or diseases such as cancer or Parkinson’s disease, for example,” says Elke Deuerling, professor of molecular microbiology at the University of Konstanz in Germany and author of the study.
The mechanism in detail
The time required for methionine cleavage and subsequent acetylation to proceed smoothly is quite short. During this time, several enzymes must be brought to the right place and regulated: MetAP1, which causes methionine cleavage, and NatA for the subsequent acetylation.
However, NatA is normally bound to an inhibitory protein, HYPK, which suppresses its function. By combining biochemical, structural and in vivo experiments, the Caltech researchers and their collaborators have now managed to shed light on how this complex process is controlled and how the macromolecules involved interact.
NAC is located at the exit of the ribosomal tunnel, where newly synthesized proteins emerge. From there, it recruits both MetAP1 and NatA and positions them with their biochemically active regions at the appropriate locations near the tunnel exit to access the nascent protein.
“In addition, NAC induces NatA to lose its inhibitory contact with HYPK. This ensures that NatA function is only activated at the ribosome, where it can then ensure the desired acetylation,” Shan explains.
Shan’s lab developed the tools that uncovered these essential roles of NAC in NatA function, providing the basis for further solving the structure of NAC bound to a ribosome with NatA and HYPK.
“It is difficult to create synchronized ribosomes with a defined nascent chain length and composition and obtain them in biophysical quantities,” Shan says. “My group has worked for many years to develop tools that allowed us to incorporate fluorescent dyes specifically into a nascent ribosome protein. This is how we detected the interaction and recruitment of these enzymes.”
Previous studies have revealed that NAC also recruits other factors into the ribosomal tunnel in addition to MetAP1 and NatA enzymes.
“We assume that the NAC has the function of an even more elaborate molecular control center,” says study co-author Martin Gamerdinger, also from the University of Konstanz. “It ensures that nascent proteins have access to different components of the cell’s biochemical toolbox when they leave the ribosome, as needed.”
The current study shows how NAC fulfills this important function in the specific case of methionine cleavage followed by acetylation. It allows scientists to better understand how deregulation of the components involved in protein modification can lead to the development of diseases. In the long term, this could serve as a basis for the development of new therapeutic approaches in medicine.
Alfred M. Lentzsch and Sowmya Chandrasekar are also co-authors of the paper “NAC guides a ribosomal multienzyme complex for nascent protein processing” at Caltech. Denis Yudin, Alain Scaiola and Nenad Ban are co-authors at ETH Zurich.
More information:
Alfred M. Lentzsch et al, NAC guides a ribosomal multienzyme complex for processing nascent proteins, Nature (2024). DOI: 10.1038/s41586-024-07846-7
Provided by California Institute of Technology
Quote:New research elucidates a key regulator of protein production (2024, August 28) retrieved August 28, 2024 from
This document is subject to copyright. Apart from any fair dealing for the purpose of private study or research, no part may be reproduced without written permission. The content is provided for informational purposes only.