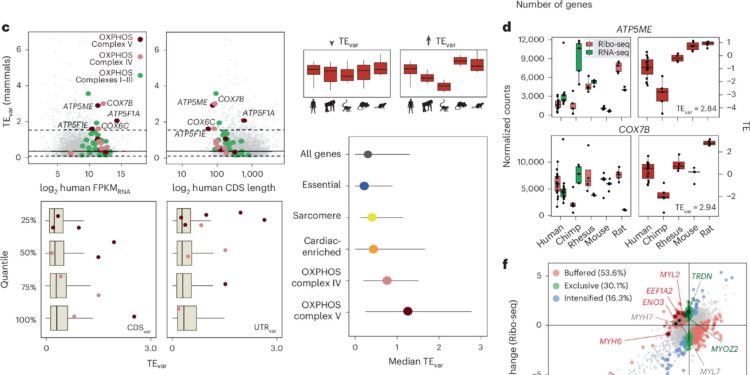
Control of translation in adult primate and mammalian hearts. Credit: Cardiovascular research on nature (2024). DOI: 10.1038/s44161-024-00544-7
A team from the Hübner and Diecke labs at the Max Delbrück Center has shown how the hearts of human and nonhuman primates differ genetically. The study, published in Cardiovascular research on naturereveals evolutionary adaptations in the human heart and provides new insights into heart disease.
Humans are 98-99% genetically similar to chimpanzees. So what explains our differences? Over the years, researchers have shown that the regulation of gene expression (when, where, and how many genes are turned on) is largely responsible for our divergent evolutionary trajectories.
Researchers from Professor Norbert Hübner’s Laboratory of Cardiovascular and Metabolic Sciences and Dr. Sebastian Diecke’s Pluripotent Stem Cell Platform at the Max Delbrück Center have uncovered surprising differences in gene expression in the hearts of humans and non-human primates. The research, led by Dr. Jorge Ruiz-Orera, highlights adaptations in the way genes are regulated that distinguish our hearts from those of our closest evolutionary relatives. It also serves as a warning against extrapolating research on animal hearts to human hearts.
“One of the most surprising findings was the dramatic difference between gene regulation in the human heart and that of other primates,” Ruiz-Orera says. In terms of anatomy, most mammalian hearts are similar. “But we have many unique evolutionary innovations in terms of gene regulation or protein translation,” he adds.
The researchers found hundreds of genes and microproteins (tiny proteins already identified in human organs but whose function remains largely mysterious) present in the human heart, but not in the hearts of other primates, rats or mice. “Many of these human genes and microproteins are also abnormally expressed in heart failure, suggesting that they may play an important role in heart function and disease and could represent new therapeutic targets,” Ruiz-Orera says.
Comparison of gene transcription and translation
The team analyzed heart tissue from chimpanzees and macaques obtained from Dr. Ivanela Kondova’s biobank at the Biomedical Primate Research Center in Rijswijk, the Netherlands, as well as stored heart tissue from humans, rats and mice that had been used in the lab’s previous research.
Using RNA sequencing, the researchers first mapped and quantified RNA molecules in heart tissue, giving them a comprehensive view of gene expression across species. To focus specifically on which regions of RNA were being translated into proteins, the researchers used Ribo-seq to sequence the RNA fragments that were being actively translated in each cell. This provided insight into which genes were producing functional proteins. By integrating data from these technologies, the team created the most comprehensive resource to date on gene and protein activity in human and non-human primate hearts.
The researchers also used cultures of cardiomyocyte cells derived from induced pluripotent stem cells (iPSC-CMs) as a model to study how genes are expressed during heart development in humans and other primates. iPSC-CMs are a useful model because they can be grown from adult primate skin cells that have been reprogrammed to an embryonic state. These cells develop into cardiomyocytes, the basic cellular unit of the heart, allowing researchers to study them at different stages of development.
Ruiz-Orera said the discovery that specific microproteins, encoded in the genome by DNA fragments called small open reading frames (ORFs), are uniquely expressed or translated in human heart cells at different stages of development, suggests that some of these genetic elements may have evolved specifically to meet the demands of the human heart. (ORFs do not have the classic characteristics of protein-coding genes and are therefore not classified as genes.)
“Our hearts have different energy needs than smaller primates like macaques, which have much faster heart rates,” he says. “This difference appears to be reflected in the regulation of genes related to energy production in the heart. These evolutionary adaptations may also be related to our bipedalism, lifestyle, and diet.”
In total, the team identified more than 1,000 species-specific genomic adaptations, including 551 genes and 504 microprotein coding regions found only in the human heart. Of these, they found 76 genes that are common to humans and other primates and mammals, but have evolved only in humans to be expressed in the heart.
Consequences for heart disease and animal use
Researchers have shown that certain human-specific genes and microproteins are deregulated in conditions such as dilated cardiomyopathy, highlighting a potential role in the development of heart disease and suggesting a new therapeutic target.
The study also raises important questions about using animals like mice to study the genetics of human heart disease. “Our results suggest that species differences can sometimes lead to misleading results,” Ruiz-Orera says. “Many genes that are expressed in the human heart are simply not expressed in the hearts of other species.”
In humans, for example, the SGLT1 gene is expressed in the heart. But in nonhuman primates, rats and mice, it is expressed only in the kidneys. Dual SGLT1 and SGLT2 inhibitors have been shown to reduce heart failure, although its exact role in the heart remains a mystery, Ruiz-Orera says. But because it is not expressed in the hearts of other animals, researchers won’t be able to learn much by testing such therapies in those models.
“This highlights the importance of taking into account the evolutionary context in medical research,” he adds.
More information:
Jorge Ruiz-Orera et al., Evolution of translational control and emergence of genes and open reading frames in the heart of human and non-human primates, Cardiovascular research on nature (2024). DOI: 10.1038/s44161-024-00544-7
Provided by the Max Delbrück Center for Molecular Medicine
Quote:Human and other primate hearts differ genetically, study finds (2024, September 24) retrieved September 24, 2024 from
This document is subject to copyright. Apart from any fair dealing for the purpose of private study or research, no part may be reproduced without written permission. The content is provided for informational purposes only.