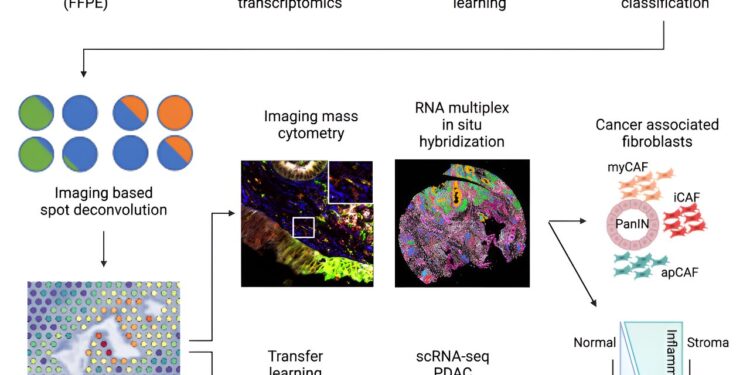
A new computational pipeline developed by Johns Hopkins Kimmel cancer researchers to identify early molecular and cellular progression of pancreatic ductal adenocarcinoma. Credit: Cellular systems (2024). DOI: 10.1016/j.cels.2024.07.001
Using a novel workflow that integrates spatial transcriptomics and machine learning for imaging analysis and integration with single-cell datasets, Johns Hopkins Kimmel Cancer Center researchers have identified novel molecular and cellular markers in the development of one of the most aggressive and deadly pancreatic cancers: pancreatic ductal adenocarcinoma (PDAC).
PDAC results from precancerous lesions in the pancreas. One type of lesion, pancreatic intraepithelial neoplasia (PanIN), can appear in the pancreas years before it develops into invasive cancer. Because PanINs are so small, they cannot be detected by conventional clinical imaging tests.
Previous analysis methods, such as bulk sequencing and single-cell sequencing, can capture the gene expression of cancer cells and other cell types in the tumor microenvironment. However, what is missing are the spatial relationships between these cell types within and around tumors, says Luciane Kagohara, Ph.D., co-senior author of the study and assistant professor of oncology at the Johns Hopkins University School of Medicine.
Elana Fertig, Ph.D., professor of oncology, director of the division of quantitative oncology sciences and co-director of the Convergence Institute and the Single-Cell Training and Analysis Center at the Johns Hopkins University School of Medicine, was also a co-senior author of the study.
Using spatial transcriptomics, a technique used to measure and map gene expression in a tissue section, the researchers developed a three-way analysis pipeline to map gene expression changes from nine patients in 14 PanINs, including five rare, high-grade PanINs. The machine learning tools used for imaging analyses (CODA) and for integration with single-cell PDAC datasets, via the innovative multiomics integration methods CoGAPS (coordinated gene activity in ensembles of motifs) and projectR, were developed in previous studies at Johns Hopkins.
The researchers made their data and code available to other researchers to uncover new insights into pancreatic cancer and, through open-source tools, to adapt spatial transcriptomics analysis methods to biomedical research in general.
The results of their work, published on August 7 in Cellular systemsprovide new insights into gene expression and spatial distribution of different cell types in the precancerous environment around PanINs. These findings are critical to understanding how PanINs evolve into PDAC, laying the foundation for future early detection of this and other types of pancreatic cancer.
Three-factor analysis revealed that some key features of pancreatic cancer were present in PanINs.
“When we looked at the progression of normal- to high-grade PanIN lesions, we found that cell proliferation progressively increases while inflammatory signaling decreases, which may be important for understanding the intrinsic low immunogenicity of these tumors,” Kagohara says. This suggests that cells in and around PanINs are already creating a more immunosuppressive environment before invasive PDAC has fully developed, she says.
The results also revealed spatial differences in the cells. “We found that cancer-associated fibroblasts, which play a major role in PDAC biology and response to treatments, were already present at the pre-malignant stage, which is consistent with previous research in animal models of pancreatic cancer but has never been observed before in humans,” Kagohara says.
“PanINs are quite small (less than a millimeter), so we were very surprised to find that even using very few cells, we were able to detect strong signatures of these lesions,” she adds.
“This is very important in the field of cancer because, for example, if we want to understand immune responses to treatments, we need to know which immune cells are closer or further away from the tumor. But we also need to understand whether there are other types of cells that block these communications by forming a physical barrier or by interacting with neoplastic cells.”
Beyond the main results, the study highlights the power of computer tools to better understand human diseases.
Spatial transcriptomics is a very powerful technology for generating data, Kagohara says, but you need the right computational tools to analyze it. “With the right tools, you can improve the results you can extract from these new technologies,” she says.
To encourage further collaboration and analysis, the study data is available on GEO (GSE254829) and the code is available on GitHub.
“As we generate more and more data on PanINs, the idea is to identify molecular signatures for the development of early detection tests,” Kagohara says. “This is one of the first studies to generate an open resource for other researchers to search for these early markers in PanINs so that we can learn more about how the spatial distribution of molecular and cellular signatures affects pancreatic cancer progression and response to treatments.”
Other co-authors of the study include Alexander Bell, Jacob Mitchell, Ashley Kiemen, Melissa Lyman, Kohei Fujikura, Jae Lee, Eron Coyne, Sarah Shin, Sushma Nagaraj, Atul Deshpande, Pei-Hsun Wu, Dimitrios Sidiropoulos, Rossin Erbe, James Chell, Lauren Ciotti, Jacquelyn Zimmerman, Denis Wirtz, Won Jin Ho, Neeha Zaidi, Elizabeth Thompson, Elizabeth Jaffee and Laura Wood of Johns Hopkins. Researchers at 10x Genomics in Pleasanton, California, contributed to this work.
More information:
Alexander TF Bell et al, PanIN and CAF transitions in pancreatic carcinogenesis revealed by spatial data integration, Cellular systems (2024). DOI: 10.1016/j.cels.2024.07.001
Provided by Johns Hopkins University
Quote:Identification of key markers of pancreatic cancer progression using a novel analysis pipeline (2024, August 29) retrieved August 29, 2024 from
This document is subject to copyright. Apart from any fair dealing for the purpose of private study or research, no part may be reproduced without written permission. The content is provided for informational purposes only.