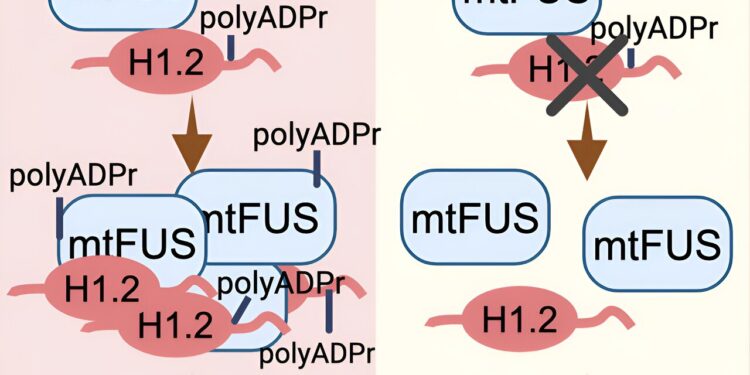
Credit: Cell Reports (2024). DOI: 10.1016/j.celrep.2024.114626
Amyotrophic lateral sclerosis (ALS) is an incurable neurodegenerative disease characterized by selective degeneration of the upper motor neurons of the motor cortex as well as the lower motor neurons of the brainstem and spinal cord.
In 90% of cases, the cause of ALS remains unknown. This is called sporadic ALS because there is no family history of the disease. There is considerable evidence that sporadic ALS may result from complex interactions between genetic predisposition and aging.
The remaining 10% of ALS cases are inherited and linked to mutations in one of 30 distinct genes involved in different cellular processes. There are severe early-onset and juvenile cases, the majority of which are caused by mutations in the FUS gene.
FUS is a protein widely expressed in tissues and plays a role in various steps of DNA and RNA processing, including DNA repair, transcription, RNA splicing, and nucleo-cytoplasmic RNA shuttling. However, mutations in this protein particularly affect motor neurons in ALS.
Professor Dr David Vilchez and his team from the CECAD Cluster of Excellence for Aging Research at the University of Cologne have identified two proteins interacting with a mutant FUS variant causing ALS (FUS P525L) by studying motor neurons derived from human induced pluripotent stem cells (iPSCs).
Their results indicate that inhibition of these interacting proteins could be a possible therapeutic target for familial cases caused by FUS mutations. The study was published under the title “ALS-FUS mutations cause abnormal PARylation and interaction with histone H1.2, leading to pathological changes” in Cell Reports.
The two proteins that interacted with the mutant FUS protein were PARP1, an enzyme that promotes poly ADP-ribosylation (PARylation), a modification that can alter proteins in different ways, and histone H1.2, a protein involved in wrapping the DNA of cells into their known chromosome shape.
In further experiments in human motor neuron cells, the researchers found that inhibiting PARylation or reducing H1.2 levels attenuated ALS-related changes such as mutant FUS protein aggregation and neurodegeneration.
The scientists then conducted experiments using the nematode Caenorhabditis elegans as a model of ALS. They found that when the worms’ homologs of the human PARP1 and H1.2 proteins were inhibited, FUS mutant aggregation and neurodegeneration also decreased.
The scientists also observed that ALS-related changes were worsened when these two proteins were overexpressed in C. elegans. “Taking all our data into account, our results indicate a link between PARylation, H1.2 and FUS with potential therapeutic implications,” said Dr. Hafiza Alirzayeva, first author of the study.
According to the researchers, the pathology between familial ALS, which was the focus of this study, and sporadic ALS is very similar: although FUS is mutated in some familial cases, non-mutant FUS also aggregates in many sporadic cases.
Professor David Vilchez, a senior researcher at CECAD, said: “Most basic research focuses on the mutant genes that cause familial ALS, because at least we know these genes. But we hope to show in further studies that our findings could also have a potential impact on sporadic ALS, because this is the form that affects the vast majority of patients.”
In future work, the authors will investigate whether these proteins might also be involved in ALS-related changes associated with other disease-causing genes, such as TDP-43 and C9orf72, as well as those in sporadic ALS.
More information:
Hafiza Alirzayeva et al, ALS-FUS mutations cause abnormal PARylation and interaction with histone H1.2, leading to pathological changes, Cell Reports (2024). DOI: 10.1016/j.celrep.2024.114626
Provided by the University of Cologne
Quote:Two proteins identified as potential targets to improve ALS symptoms (2024, August 22) retrieved August 22, 2024 from
This document is subject to copyright. Apart from any fair dealing for the purpose of private study or research, no part may be reproduced without written permission. The content is provided for informational purposes only.